
In this tutorial I will take you through how to:
- Read in data
- Perform feature engineering, dummy encoding and feature selection
- Splitting data
- Training an XGBoost classifier
- Pickling your model and data to be consumed in an evaluation script
- Evaluating your model with Confusion Matrices and Classification reports in Sci-kit Learn
- Working with the shap package to visualise global and local feature importance
Before we get going I must explain what Shapley values are?
A short primer on Shapley Values
Shapley values were created by Lloyd Shapley an economist and contributor to a field called Game Theory. This type of technique emerged from that field and has been widely used in complex non-linear models to explain the impact of variables on the Y dependent variable, or y-hat.
General idea
General idea linked to our example:
You have trained a machine learning model to predict whether a patient will be stranded or not. A stranded patient is an NHS England term for a patient that has been in an inpatient setting for 7 consecutive days or more. We now need to explain the prediction. The patients who are stranded normally have other issues, such as ageing, care home residency, etc. We use previous class labels and membership to work out the effect of each feature i.e. the weight of the feature and its strength on the classification, for regression it would be how much it contributes on average to the dependent variable.
Shapley metrics come into play with payouts and the contribution of the feature to that payout, so for classification models, it would be how likely they are to cause a classification/outcome over other features. The goal with classification would be to explain the difference between someone who is classified as a stranded patient over those that are not stranded.
The terms players, game and payout are used in the literature and refer to the game as the prediction task for a single instance on that dataset, i.e. one patient observation. The gain is the actual prediction/classification label for that observation minus all the other observations, in a classification sense, it would be probabilities. The players are the unique features in our model that interact in the game to achieve some loss or gain.
Simply then, this is repeated for all observations in the data and the predictions averaged for regression over all the marginal contributions and possible coalitions. These could be the possible coalitions:
- No feature values
- Age of patient
- Previous times entered care
- Care home resident
- Age of patient + Previous times entered care
- Age of patient + Care home resident
- And various other permutations
For each of these coalitions, we compute the predicted class label with and without the feature value age of the patient and take the difference to get the marginal contribution. The Shapley value is the (weighted) average of marginal contributions. We replace the feature values of features that are not in a coalition with random feature values from the stranded patient dataset to get a prediction from the machine learning model.
For those interested in reading more check out the Wikipedia page, which has the in-depth workings of how the Shapley Values are worked through, but the master equation is presented hereunder:
where n is the total number of players and the sum extends over all subsets S of N not containing player i. The formula can be interpreted as follows: imagine the coalition being formed one actor at a time, with each actor demanding their contribution 

This might not make sense yet, but it soon will become apparent, plus let’s get coding in Python.
Building the model to test out on the Shap package
The following code will import the relevant packages needed to work through this tutorial.
Data Import and Package Initialisation
# =============================================================================
# Import necessary packages
# =============================================================================
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
from xgboost.sklearn import XGBClassifier
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import LabelBinarizer
# =============================================================================
# Data reading and cleansing
# =============================================================================
# Read in data
df = pd.read_csv("https://hutsons-hacks.info/wp-content/uploads/2021/04/LongLOSData.csv")
df = df.dropna()
# Encode categorical variables in X data frame
Y = df["stranded.label"]
# Select df columns for X
drop_cols = ["stranded.label", "admit_date"]
X = df.loc[:, [x for x in df.columns if x not in drop_cols]]
# Dummy encode the categorical labels
X = pd.get_dummies(X, columns=['frailty_index'])
# Drop one reference column as it does not give us much information and could
# cause multicollinearity affects in linear models
X = X.drop(["frailty_index_No index item"], axis=1)
Taking your through the data preparation process:
- Use pandas to read in a csv from this website, based off the NHSRDatasets package that contains the stranded_data file.
- Next I use df.dropna() to get rid of NA values in the data frame
- I then use list comprehension to drop the columns in the drop_cols variable and update the X variable – this being where all my indepdent variables are stored
- The final step is to dummy encode or one-hot encode the categorical variables using the get_dummies pandas function to encode the relevant columns. These columns I pass as a list [].
Feature Selection
The next step is to drop the variables that have little impact on the prediction. I do this as a step to see which variable it eliminates automatically, but I will retrain them for the shap value generation part of the process.
The following code implements a function to perform the recursive feature elimination and then we use a fit object to remove these from a variable:
# =============================================================================
# Feature Selection using Recursive Feature Engineering
# =============================================================================
# Feature selection using RFE
def recursive_feature_eng(model, X, Y):
print("[INFO] Starting Recursive Feature Engineering")
rfe = RFE(model)
rfe_fit = rfe.fit(X,Y)
print("Number of features chosen: %d" % rfe_fit.n_features_)
print("Selected features chosen: %s" % rfe_fit.support_)
print("Fit ranking of feature importance: %s" % rfe_fit.ranking_)
print("[INFO] Ending Recursive Feature Engineering")
return [rfe_fit, rfe_fit.n_features_, rfe_fit.support_]
rfe_model = LogisticRegression(solver='liblinear')
rfe_fit = recursive_feature_eng(rfe_model, X, Y)
# Pull out the feature ranking from the fitted object
columns_to_remove = rfe_fit[2]
X_reduced = X.loc[:,columns_to_remove]
To understand the steps here:
- We use a base model, for this it is logistic regression
- We then fit using the fit object from sci-kit learn
- The fit extracts a number of features, these are:
- n_features_ – this highlights the number of features selected by the elimination algorithm
- support_ – this extracts a boolean array containing False and True – the True values are those to be retained in the data, with False showing those features to be removed
- ranking_ – this is the variable importance ranking using the selected model
- This function returns these items in a list
In the next couple of steps, we fit the model, utilise our custom function, specify in the columns_to_remove variable that we want to extract the list item at index 2, which is the boolean identity array and finally, x_reduced uses this identity array to select only the True columns from the feature selection algorithm.
Outputs of the function in the console window are as below:
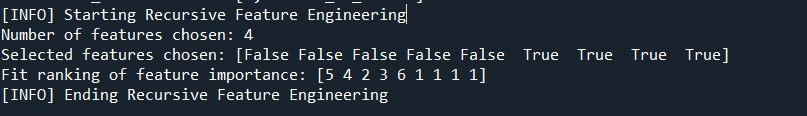
Hold out splitting the data and transforming stranded Y label
Here I will separate the data into training and testing partitions, as well as binarising the prediction column:
# ============================================================================= # Transform Y and Split the data # ============================================================================= # Transform Y label lb = LabelBinarizer() Y = lb.fit_transform(Y) #Split the data X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=.3, stratify=Y)
The LabelBinarizer() converts categorical strings into an encoding of 0 or 1, as this is a binary classification problem. Thus, the encodings for Not Stranded = 0 and Stranded = 1.
Train and test splitting creates four variables X_train, X_test, Y_train and Y_test and splits them by a proportion of 70% into the ML model training sample and 30% in the validation testing sample. We will use this, later on, to test the accuracy of the model on supposedly unseen data, as eventually, we would like to put this model into production. Finally, stratify does a stratified split on the class labels to try and make sure that the labels are not imbalanced in terms of class representation i.e. more patients from the stranded class than the not stranded class.
Training the XGBoost Model
The penultimate step is to train the model on a training set to make predictions based on all the historic observations of the probability of being a stranded patient or not stranded. We implement this below:
# =============================================================================
# Train the model
# =============================================================================
model = XGBClassifier(n_estimators=1000, max_depth=10, learning_rate=0.001,
use_label_encoder=False, eval_metric='logloss')
xgb_fit = model.fit(X_train, Y_train)
Here I used some random known hyperparameter settings. The n_estimators is the number of trees, the depth of the tree is max_depth in terms of the number of nodes to branch down to, the learning rate is set to a known good parameter, albeit all these can be tuned. Additionally, I have had to set a couple of extra options to stop warnings from appearing, but they are more about the internals of the package I used.
Pickling the model
Finally, I am going to pickle the model so that I can use it in the evaluation and feature importance script:
# ============================================================================= # Pickle model to work with it in next script # ============================================================================= from pickle import dump, load filename = "stranded_model.sav" dump([model, X_train, X_test, Y_train, Y_test], open(filename, 'wb'))
This sets a filename for the model and uses the dump method to save, as a list structure, the model and training data. I then use the open method to write this to binary (‘wb’).
To confirm this has been written I switch over to my File Explorer and see if it is in my working directory:

We now have everything we need to consume the model in the next tutorial.
The training script for the first half of this can be found hereunder:

Model Evaluation and Global / Local Feature Importance with the Shap package
The steps now are to:
- Load our pickle objects
- Make predictions on the model
- Assess these predictions with a classification report and confusion matrix
- Create Global Shapley explanations and visuals
- Create Local Interpretability of the Shapley values
Loading Pickle objects and unpacking
import shap from pickle import load import matplotlib.pyplot as plt import numpy as np # ============================================================================= # Load in pickle file # ============================================================================= filename = "models/stranded_model.sav" # Previous pickle file returned a list so we will perform # multiple assignment here model, X_train, X_test, Y_train, Y_test = load(open(filename, 'rb'))
A couple of things to point out here. I load the files from the pickle file in the models’ directory and the model package name stranded_model.sav. I then do multiple assignment on the contents, as the file contains the trained model and the testing files. Once completing this you should have five variables in memory allocated to the model, X_train, X_test, Y_train and Y_test variables, as well as a string variable for the filename, or better put path.
Making predictions and evaluating
I will now make predictions on the data and store these in two variables i.e. pred_class and pred_probs. Printing these out would show a) the predicted class labels i.e. stranded or not and b) the probabilities of being in class A over class B. This is implemented below:
# ============================================================================= # Make model predictions # ============================================================================= # Make predictions with model pred_class = model.predict(X_test) pred_probs = model.predict_proba(X_test)
Now I have made predictions I can evaluate the model based on the ground truth labels in the testing set vs the predictions I have just made. I have created a function to help with this:
def confusion_matrix_eval(Y_truth, Y_pred):
from sklearn.metrics import confusion_matrix, classification_report
cm = confusion_matrix(Y_test, pred_class)
cr = classification_report(Y_test, pred_class)
print("-"*90)
print("[CLASS_REPORT] printing classification report to console")
print("-"*90)
print(cr)
print("-"*90)
return [cm, cr]
cm = confusion_matrix_eval(Y_test, pred_class)
Printing out the cm object I get the classification report formatted as in the function:
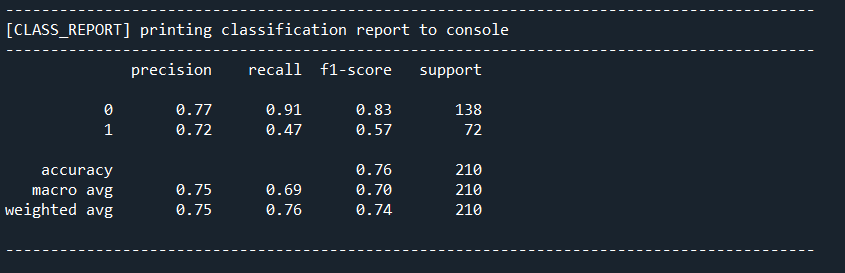
The overall accuracy is not bad. To explain what the other metrics mean:
- Precision – out of the positive predicted what are truly positives = Precision = TP / (TP+FP)
- Recall – out of the total positive, what percentage are predicted positive, same as the true positive rate = Recall = TP / TP + FN
- F1 score = 2 * (Precision * Recall)/(Precision + Recall) – this is the harmonic mean of the models precision and recall
This shows that generally the model is good at predicting and is relatively balanced, however it is more precise at predicting those patients who are not stranded.
Working with Shap for Global and Local Interpretability
I will now create an explainer object and specify the shap values. The explainer is trained on the model and the shap_values are a method attached to that. To implement we will use the TreeExplainer method, although there are a number of other explainers for deep learning, computer vision and NLP.
# ============================================================================= # Shapley Values for Feature Importance # ============================================================================= # Fit relevant explainer explainer = shap.TreeExplainer(model) shap_values = explainer.shap_values(X_train) # View shap values print(shap_values)
The shap values represent the relative strength of the variable on the outcome and it returns an array, I have implemented a print statement to observe this:
Printing the shape of the array you should see that it should contain the same amount of rows and columns as your training set. Mine returns a tuple of (489, 9).
Global Importance
To obtains a global importance plot of the effects of the features on whether a patient is stranded the shap package has a summary_plot function, this can be implemented like so:
# =============================================================================
# # Get global variable importance plot
# =============================================================================
plt_shap = shap.summary_plot(shap_values, #Use Shap values array
features=X_train, # Use training set features
feature_names=X_train.columns, #Use column names
show=False, #Set to false to output to folder
plot_size=(30,15)) # Change plot size
# Save my figure to a directory
plt.savefig("plots/global_shap.png")
This produces a Shapley value chart to allow us to look at the affects of the relevant features on whether a patient stays in hospital longer than 7 days, otherwise known as a stranded patient:
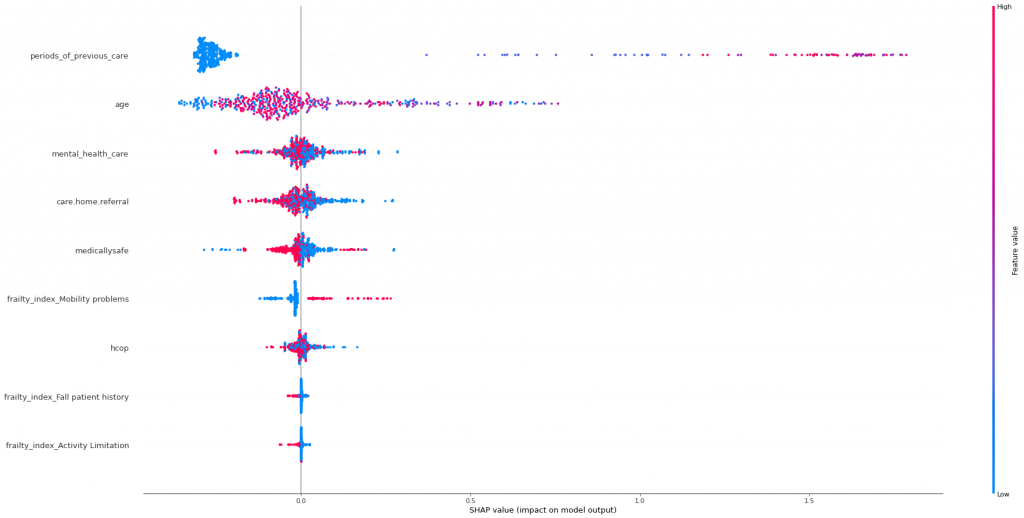
From the chart you can see:
- Periods of previous hospital care show a massive effect on whether a patient will be stranded, as most gravitate to the positive plane and show a marked distribution
- Age has more of an effect on some patients than other, so this would be best viewed from a local perspective.
- Mental health care seems to have an even split
Please bear in mind this dataset is only for demo purposes and does not represent actual hospital activity.
I like this, but I would like to understand what drives it on a patient by patient level. That is where local interpretability come in.
Local Importance
This works on a patient by patient-level or observation by observation level. I will create an obs_idx variable to store the integer location of a patient in the data, for the sake of example. This plot is implemented very similar to the above:
# =============================================================================
# # Local Interpretation Plots
# =============================================================================
obs_idx = 488 # Relates to a specific patient observation
local_plot = shap.force_plot(explainer.expected_value,
shap_values[obs_idx],
features=X_train.loc[obs_idx],
feature_names=X_train.columns,
show=False, matplotlib=True)
# Save my plot
plt.savefig("force_plot.png")
Viewing patient record 488 we get the following:
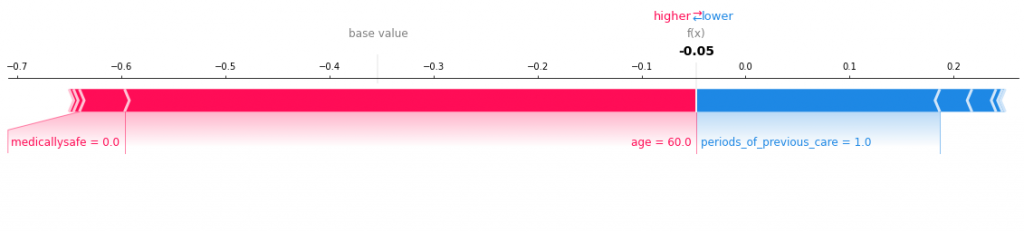
This show age is a bigger driver than anything else. Let’s try another one – say patient 3:
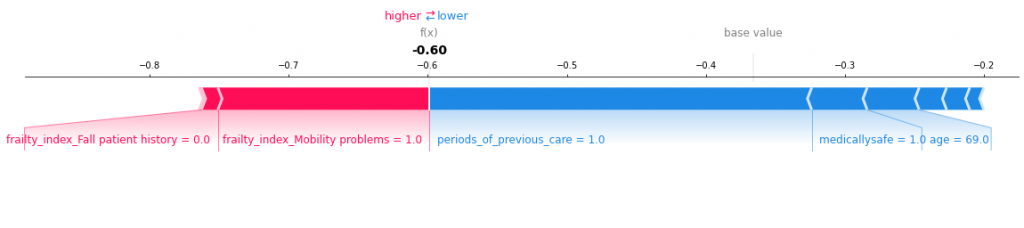
A more mixed bag this time, as period of previous care drive the function lower, whereas frailty issues have a higher effect. Patient 10 looks like below:
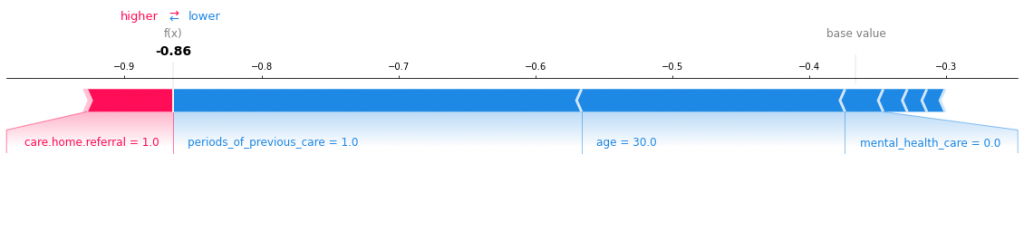
I like these plots as it allows us to get to the bottom of what is going off per patient and what might be affecting whether they are stranded. This could aid clinicians in providing tailored treatment to each patient based off these attributes. The code for this workbook is here:

Conclusion
We have learned a lot here today. We started with an ML classification task, then we pickled and saved our model and data. Followed by loading this and then evaluating the model fit and working with the shap package.
Time for a cup of tea and relax. If you made it to the end, you deserve one!

