Thanks to the NHS-R community I have had time to work on another package, due to their pledge to get more packages in R funded. A big thanks to Mohammed Amin Mohammed and all the R community team.
This package utilises all the excellent lookups provided by NHS Digital and the NHS Data Dictionary and allows you to access these lookups in one place.
What problem does this solve?
Many times I have worked with trusts and they have to download these lookups and build them in their data warehouse. The problem is this requires local management of the databases to make sure they are up to date and in one place.
This package aims to centralise and standardise this method, so every healthcare agency can use the same lookups across the UK.
Additionally, this package found its way to RStudios Top 40 packages to watch out for in January 2021: https://rviews.rstudio.com/2021/02/24/january-2020-top-40-new-cran-packages/.
When does the package launch?
The package launches officially in April 2021 and the NHS-R community are holding a launch webinar on the 21st April 2021. To view this navigate to webinars on the NHSR community web page.
The recording of this webinar will be made available after the webinar and I will follow up with a subsequent post detailing what I have learned from my first CRAN submitted package, and why I normally just stick them on GitHub to be downloaded.
How many hits has the package had?
Utilising the dlstats package, see associated post, the package has already had a number of downloads:
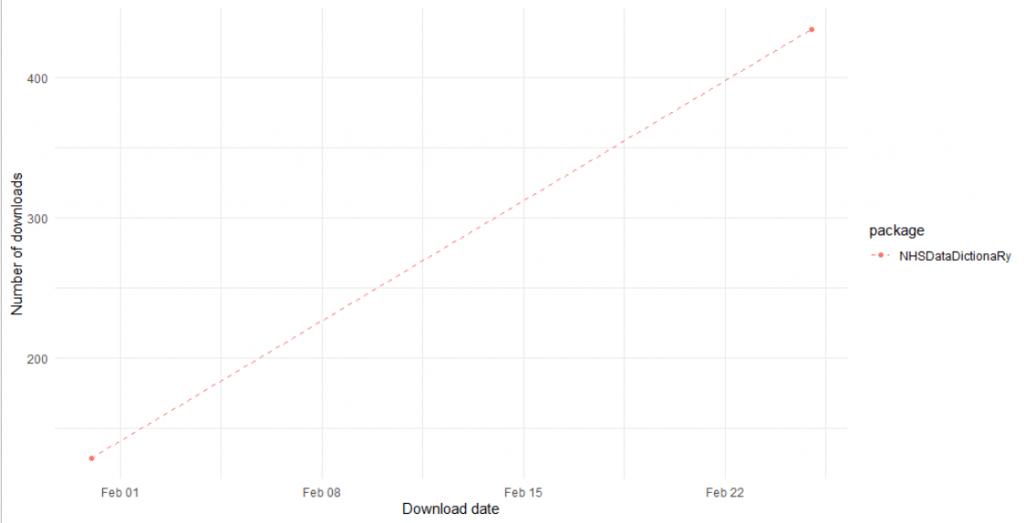
This shows that I have about 563 downloads of the package from CRAN, and this has not yet been launched officially. Not a bad couple of days work!
But, what is this package and how might it help you if you work for the NHS and work in R. The following section will detail this.
What the package does?
The vignette attached to this package explains how to use the package and what it is for.
In essence, the package contains:
- nhs_data_elements() function – this function will return all the current data element lookups from the NHS Data Dictionary and returns these as a tibble. This acts as the master lookup for all of the other functions that are contained in the package
- Text manipulation convenience functions for old Excel users:
- left_xl() – performs a left trim on a character string
- right_xl() – performs a right trim on a character string
- mid_xl() – performs a middle text extraction
- len_xl() – a simple wrapper to return the number of characters in a string
- Getting all the current hyperlinks on a page, stored in a tibble. The function for this is linkScrapeR
- TableR and scrapeR are the two powerhouses of the package and can be utilised alongside the nhs_data_element() function to extract a lookup and then use this lookup alongside existing NHS data:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 | # Filter by a specific lookup required reduced_tibble <- dplyr::filter(nhs_tibble, link_name == "ACTIVITY TREATMENT FUNCTION CODE") #Use the tableR function to query the NHS Data Dictionary website and return the associate tibble treatment_function_lookup <- NHSDataDictionaRy::tableR(url=reduced_tibble$full_url, xpath = reduced_tibble$xpath_national_code, title = "NHS Hospital Activity Treatment Function Codes") # The query has returned results, if the url does not have a lookup table an error will be thrown print(head(treatment_function_lookup,10)) #> # A tibble: 10 x 4 #> Code Description Dict_Type DttmExtracted #> <chr> <chr> <chr> <dttm> #> 1 199 Non-UK provider; TREATMENT FU~ NHS Hospital Activi~ 2021-01-14 17:10:08 #> 2 499 Non-UK provider; TREATMENT FU~ NHS Hospital Activi~ 2021-01-14 17:10:08 #> 3 100 General Surgery Service NHS Hospital Activi~ 2021-01-14 17:10:08 #> 4 101 Urology Service NHS Hospital Activi~ 2021-01-14 17:10:08 #> 5 102 Transplant Surgery Service NHS Hospital Activi~ 2021-01-14 17:10:08 #> 6 103 Breast Surgery Service NHS Hospital Activi~ 2021-01-14 17:10:08 #> 7 104 Colorectal Surgery Service NHS Hospital Activi~ 2021-01-14 17:10:08 #> 8 105 Hepatobiliary and Pancreatic ~ NHS Hospital Activi~ 2021-01-14 17:10:08 #> 9 106 Upper Gastrointestinal Surger~ NHS Hospital Activi~ 2021-01-14 17:10:08 #> 10 107 Vascular Surgery Service NHS Hospital Activi~ 2021-01-14 17:10:08 act_aggregations <- tibble(SpecCode = as.character(c(101,102,103, 104, 105)), ActivityCounts = round(rnorm(5,250,3),0), Month = rep("May", 5)) # Use dplyr to join the NHS activity by specialty code act_aggregations %>% left_join(treatment_function_lookup, by = c("SpecCode"="Code")) #> # A tibble: 5 x 6 #> SpecCode ActivityCounts Month Description Dict_Type DttmExtracted #> <chr> <dbl> <chr> <chr> <chr> <dttm> #> 1 101 251 May Urology Service NHS Hospita~ 2021-01-14 17:10:08 #> 2 102 250 May Transplant Sur~ NHS Hospita~ 2021-01-14 17:10:08 #> 3 103 248 May Breast Surgery~ NHS Hospita~ 2021-01-14 17:10:08 #> 4 104 247 May Colorectal Sur~ NHS Hospita~ 2021-01-14 17:10:08 #> 5 105 248 May Hepatobiliary ~ NHS Hospita~ 2021-01-14 17:10:08 # This easily joins the lookup on to your data |
Further details of how to use all the functions can be found in the supporting package vignette.
Click GitHub to download the GitHub version of the package.
Credits
I would like to say thank you to my organisation (Arden and GEM CSU) for allowing the development of this package to take place, especially my line manager Jess Hicks and to Mohammed A Mohammed, the lead for the NHS-R Community for giving me the grant to undertake this work.
I look forward to making more developments of this, hopefully useful, package in the future.

Comments are closed.