I had a recent enquiry on our NHS-R community slack channel about which package is best to work with larger datasets, such as 5 million plus records, with high dimensionality. I got to thinking that the DBI and odbc packages would be best for this purpose, as this is still not considered big data, it is more medium data with manageable dimensionality.
The functions can be fiddly to understand, so I thought I would combine all the functions into one lovely function wrapper in R.
Overview of the function
The function below allows you to pass in your connection parameters, for SQL Server trusted connections (I can extend this if you need to specify user name and password for SQL Server databases). The function is made up of below:
Stepping through the function we have:
- Input parameter:
- SQL Server driver – the driver to connect the function to the database. The connection strings website should help you to navigate between these functions.
- SQL Server server – the name of the server to connect to
- SQL Server database (db) i.e. the database you want to connect to
- SQL Server schema – the default schema is dbo and is defaulted, but this allows you to save the tables against any table schema
- The R (tbl) object – normally a data.frame or a tibble to pass into it
- Append – this is defaulted to FALSE, but to make the query an append query switch this to TRUE and the overwrite function to FALSE
- Overwrite – set to TRUE by default, meaning a new table will be created everytime the function is run
- tbl_name – the name to give the table in SQL server
- Input params (params) – this essentially saves everything that the user enters into the function as a list
- Connection object (con) – this passes the inputs into the dbConnect function from the odbc package
- Timings – a system.time wrapper is wrapped around the odbc::dbWriteTable function to time how long the table takes to write to the SQL database
- The parameters list gets updated with the timings and returned to the user
That is the entirety of the function, but this saves on retyping and having to instantiate the connection object before saving to the database.
Using the write_table_sql() function
The function has been created. What I am going to do now is load in the brilliant NHSRdatasets package built by my friend Chris Mainey and we are going to load the ons_mortality() data package. This contains deaths at an aggregate level:
I have the relevant dataset. I need to store this in my SQL server database, using a trusted connection. This is how simple it is now with the wrapper function:
Here I pass in:
- The SQL drive
- The server name – this will vary dependent on your installation
- The database
- The schema, as stated, this defaults to dbo, if custom schemas have not been created in SQL
- The tbl – which is the R data.frame or tibble object
- Append = TRUE
- Overwrite is set to FALSE
- The tbl_name is the table for the SQL server table
Running this, the following is returned:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 | Establishing connection to: localhost\SQLEXPRESS Writing to database - please wait... Finished writing to database: RDatabase in 0.63 seconds. > print(mort_sql_list) $input_params $input_params$driver [1] "SQL Server" $input_params$server [1] "localhost\\SQLEXPRESS" $input_params$database [1] "RDatabase" $input_params$trusted [1] "True" $input_params$db_schema [1] "data" $input_params$db_tbl # A tibble: 18,803 x 5 category_1 category_2 counts date week_no <chr> <chr> <dbl> <date> <int> 1 Total deaths all ages 12968 2010-01-08 1 2 Total deaths all ages 12541 2010-01-15 2 3 Total deaths all ages 11762 2010-01-22 3 4 Total deaths all ages 11056 2010-01-29 4 5 Total deaths all ages 10524 2010-02-05 5 6 Total deaths all ages 10117 2010-02-12 6 7 Total deaths all ages 10102 2010-02-19 7 8 Total deaths all ages 10295 2010-02-26 8 9 Total deaths all ages 9981 2010-03-05 9 10 Total deaths all ages 9792 2010-03-12 10 # ... with 18,793 more rows $input_params$append_bool [1] TRUE $input_params$overwrite_bool [1] FALSE $input_params$table_name [1] "onsMortality" $run_time [1] 0.63 |
Accessing individual parameters from the function (params)
To access individual parameters you use the dollar notation to access fields:
Checking the data has made it to SQL Server
I navigate to my SQL Server instance, set on my localhost in this instance, and can find the data under the relevant table name onsMortality:
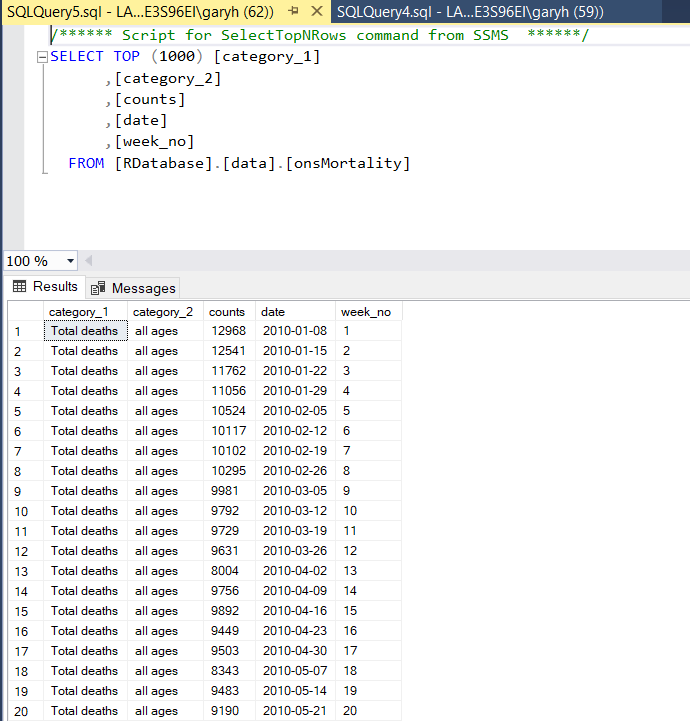
As you can see the R function has worked. What is great is that I tested this with 50 million records and 12 columns and the DBI function took 24 minutes, whereas with RODBC this take 1 hour plus.
Conclusion
This function is ready to be dropped into your projects and please have a play to add the functionality to the function to get it to work with none trusted connections i.e. those that require a user ID and password.
I hope you find this function useful, and it does really speed the process up.
To find this go to my GitHub page.

